The Brain Functional Genome Program (BFGP) aims to deepen our understanding of the human brain during development by linking gene expression locations to their functions, providing the broad community working on neurological and neurodegenerative diseases with established and robust techniques and targeted tools. It is based on cutting‑edge technologies in which Paris Brain Institute is an expert: 3D visualization (iDISCO) of transcripts, spatial transcriptomics, and the generation of induced pluripotent stem cell (iPSC) lines.
The Brain Functional Genomics Project (BFGP) is a multidisciplinary initiative at the Paris Brain Institute that aims to build an integrated resource linking spatial gene expression, protein distribution, and gene function in the developing brain.
The project is organized around three complementary aims.
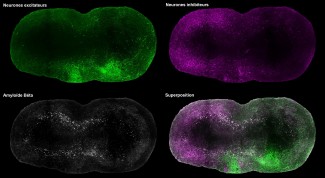
- The first aim develops a novel workflow combining 3D RNA fluorescence in situ hybridization (3D RNA FISH) with spatial transcriptomics (Visium HD, Visium V2, GeoMx) and spatial proteomics on the same tissue samples. Applied to human fetal and mouse brain tissue at key developmental stages, this approach will generate a spatially resolved, multimodal atlas of gene expression during brain development.
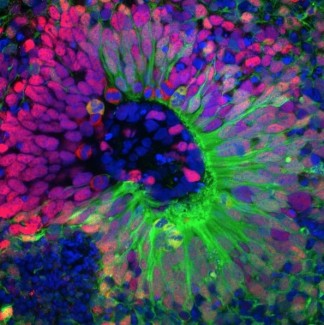
- The second aim establishes a functional screening platform based on CRISPRi-mediated gene knockdown in iPSC-derived cortical organoids. Using inducible and constitutive dCas9 systems, the screen will systematically assess the impact of disease-associated gene downregulation on neurogenesis, providing a direct link between gene function and neurodevelopmental phenotypes.
- The third aim develops the computational infrastructure required to integrate these datasets, including pipelines for 3D-to-3D and 3D-to-2D image alignment, spatial transcriptomics analysis at single-cell resolution, and metadata databases ensuring full traceability from sample to data.
Together, these three aims will produce the first atlas connecting spatial gene expression patterns with functional gene roles in the developing brain. The project leverages unique technological capabilities at the institute and fosters cross-disciplinary collaboration between experimental biologists, engineers, and data scientists. All tools, pipelines, and datasets will be made available to the research community under open science principles.
Ambition
Understanding how the brain develops and how this process goes awry in neurological and psychiatric diseases requires more than cataloguing which genes are expressed where. It demands an integrated view — one that captures gene expression in its native three-dimensional tissue context, links it to protein distribution, and connects spatial patterns to gene function. This is precisely the ambition of the Brain Functional Genomics Project. By combining, for the first time on the same samples, 3D RNA FISH with spatial transcriptomics and spatial proteomics, and by coupling this multimodal atlas with systematic functional screening in human cortical organoids, BFGP will produce a resource that bridges the gap between spatial gene expression and biological function during brain development. No existing atlas offers this level of integration. The computational tools and analysis pipelines developed within the project will make this resource fully accessible and reproducible, establishing an open platform for the neuroscience community. Ultimately, by revealing where, when, and in which cell types disease-associated genes act — and what happens when they are disrupted — BFGP will lay the groundwork for understanding the cellular mechanisms through which genetic risk translates into neurological disease, and for identifying the cell populations and developmental windows most relevant to future therapeutic strategies.
AIM-1: A new paradigm for multimodal spatial profiling of intact tissues.
Spatial transcriptomics technologies have transformed our understanding of RNA distribution within tissues, offering unprecedented insights into cellular organization and tissue architecture. However, current approaches remain largely confined to thin two-dimensional sections, limiting our ability to capture the complex three-dimensional structure of organs such as the brain. Our project addresses this fundamental limitation. The workflow developed in AIM-1 is, to our knowledge, the first protocol enabling the detection of multiple RNA targets in three dimensions by 3D RNA FISH (six genes across two hybridization rounds), followed by spatial transcriptomics (Visium HD, Visium V2, or GeoMx) and spatial proteomics (up to 20 antibodies) — all performed sequentially on the same sample. This multimodal, same-sample strategy is particularly critical for rare and irreplaceable specimens, such as human fetal brain tissue or patient-derived tumor samples, where every section must yield maximum information. The method paper currently in preparation — covering both the experimental protocol and the associated analysis pipeline — is expected to have a significant impact across multiple fields, including developmental neuroscience, neuro-oncology, and any discipline requiring spatially resolved molecular profiling of complex tissues. Beyond neuroscience, the workflow is in principle applicable to any organ or tissue type, and its open dissemination will enable adoption by other laboratories worldwide.
AIM-2: An accessible functional screening platform for neurodevelopmental biology.
Recent years have seen significant progress in large-scale functional screening in three-dimensional organoid models, primarily through pooled approaches in which thousands of genetic perturbations are introduced simultaneously and read out by single-cell sequencing (e.g., CROP-seq, Perturb-seq in organoids). While powerful, pooled screens are limited in the phenotypic dimensions they can capture: readouts are typically restricted to transcriptomic changes, and spatial or morphological phenotypes — which are central to understanding brain development — are largely inaccessible. Our project takes a fundamentally different approach by developing an arrayed CRISPRi screening platform in iPSC-derived cortical organoids, in which each gene knockdown is performed and phenotyped individually. This arrayed format enables the assessment of rich, multi-dimensional phenotypes including morphology, cellular composition, spatial organization, and neuronal activity — none of which can be readily captured in a pooled setting. To our knowledge, this is the first effort to establish a systematic arrayed screen at scale in cortical organoids. The development of both an inducible and a constitutive all-in-one lentiviral system provides flexible delivery tools compatible with arrayed experimental designs. The extensive troubleshooting and optimization conducted during this period — including resolution of Cre leakiness and GCaMP detection issues — represent practical knowledge that will benefit any group attempting similar approaches. These tools and protocols will provide the community with an accessible framework for arrayed functional genomics in organoid systems, filling a critical gap between pooled screening approaches and detailed mechanistic studies. In the longer term, the ability to phenotype individual gene knockdowns in a human-relevant three-dimensional model opens the door to identifying not only disease-relevant genes but also the specific cellular and structural phenotypes through which they act — information that is essential for prioritizing therapeutic targets.
AIM-3: Open computational infrastructure for multimodal data integration.
The integration of heterogeneous spatial datasets — 3D volumetric imaging, 2D spatial transcriptomics, spatial proteomics — poses computational challenges that no existing off-the-shelf software suite fully addresses. The pipelines developed in AIM-3 for 3D-to-3D image registration, 3D-to-2D alignment, and nuclear segmentation-based spatial single-cell analysis represent novel contributions to the bioimage informatics and spatial omics analysis communities. These tools are being developed with open release in mind, ensuring reproducibility and broad accessibility. The metadata databases designed for sample-to-data traceability also address a growing need in the field for rigorous data management in complex, multi-step experimental workflows — a prerequisite for reproducible science and future data sharing under FAIR principles. These contributions extend beyond our specific biological questions and will be relevant to any group working with multimodal spatial data, in neuroscience or other biomedical disciplines.
Taken together, the three aims of BFGP contribute an integrated technological framework — from tissue to data to biological insight — that did not previously exist. By making all protocols, tools, and datasets openly available, the project is designed to serve as a community resource rather than a closed platform. The combination of spatial atlas data with functional screening results has the potential to reshape how the neuroscience community prioritizes candidate disease genes, by providing not only expression maps but direct evidence of gene function in a human-relevant developmental model. This approach is transferable to other organ systems and disease contexts, and we anticipate that the methodological contributions of BFGP will have a lasting impact on how multimodal spatial biology experiments are designed, executed, and analyzed.